Chemical Industry and Engineering Progress ›› 2021, Vol. 40 ›› Issue (3): 1202-1214.DOI: 10.16085/j.issn.1000-6613.2020-2101
• Special column:Green biomanufacturing • Previous Articles Next Articles
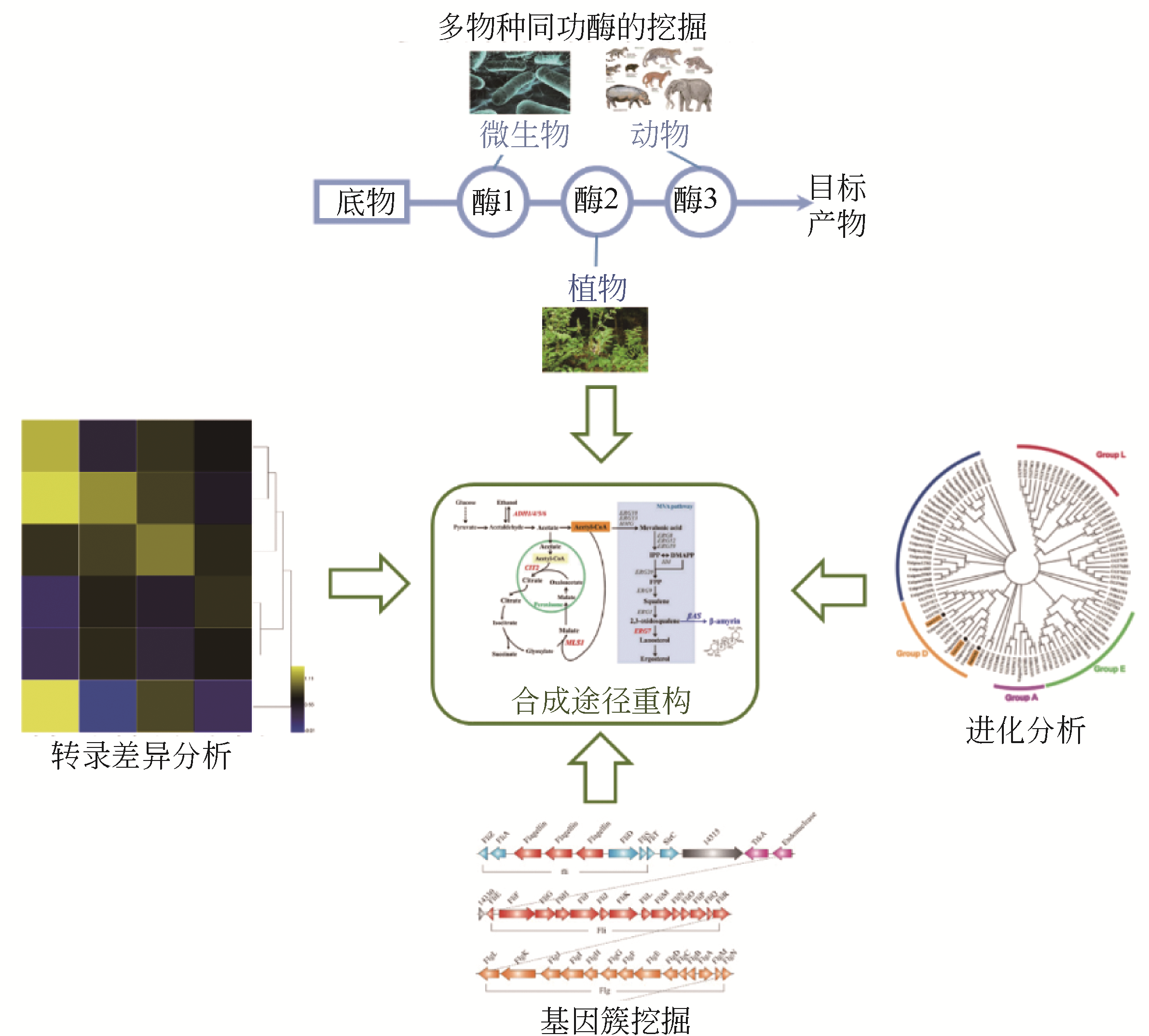
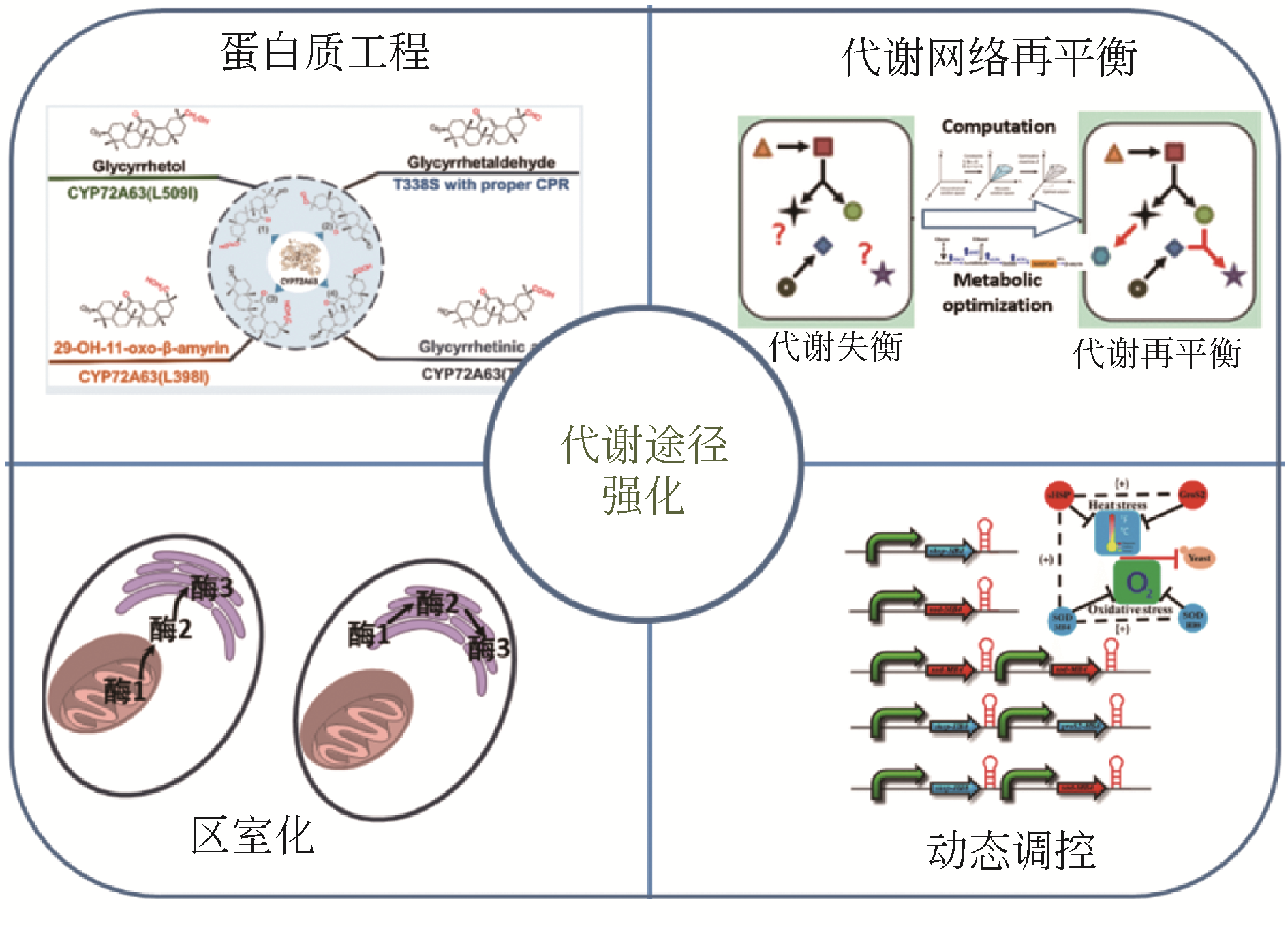
Design and construction of microbial cell factory for biosynthesis of plant natural products
- Department of Chemical Engineering, Tsinghua University, Beijing 100084, China
-
Received:2020-10-20Online:2021-03-17Published:2021-03-05 -
Contact:LI Chun
微生物合成植物天然产物的细胞工厂设计与构建
- 清华大学化学工程系,北京 100084
-
通讯作者:李春 -
作者简介:孙文涛(1988—),男,博士,研究方向为代谢工程与合成生物学。E-mail:sunwentao2016@126.com 。 -
基金资助:国家重点研发计划(2018YFA0901800);国家自然科学基金(21736002)
CLC Number:
Cite this article
SUN Wentao, LI Chun. Design and construction of microbial cell factory for biosynthesis of plant natural products[J]. Chemical Industry and Engineering Progress, 2021, 40(3): 1202-1214.
孙文涛, 李春. 微生物合成植物天然产物的细胞工厂设计与构建[J]. 化工进展, 2021, 40(3): 1202-1214.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: https://hgjz.cip.com.cn/EN/10.16085/j.issn.1000-6613.2020-2101
| 1 | KRIVORUCHKO A, NIELSEN J. Production of natural products through metabolic engineering of Saccharomyces cerevisiae[J]. Current Opinion in Biotechnology, 2015, 35: 7-15. |
| 2 | CHEMLER J A, KOFFAS M A. Metabolic engineering for plant natural product biosynthesis in microbes[J]. Current Opinion in Biotechnology, 2008, 19(6): 597-605. |
| 3 | ZHANG G L, CAO Q, LIU J Z, et al. Refactoring β-amyrin synthesis in Saccharomyces cerevisiae[J]. AIChE Journal, 2015, 61(10): 3172-3179. |
| 4 | PANDEY R P, PARAJULI P, KOFFAS M A G, et al. Microbial production of natural and non-natural flavonoids: pathway engineering, directed evolution and systems/synthetic biology[J]. Biotechnology Advances, 2016, 34(5): 634-662. |
| 5 | YU Y, CHANG P C, YU H, et al. Productive amyrin synthases for efficient alpha-amyrin synthesis in engineered Saccharomyces cerevisiae[J]. ACS Synthetic Biology, 2018, 7(10): 2391-2402. |
| 6 | SUN W T, QIN L, XUE H J, et al. Novel trends for producing plant triterpenoids in yeast[J]. Critical Reviews in Biotechnology, 2019, 39(5): 618-632. |
| 7 | XU Y H, LIU Y L, RASOOL A, et al. Sequence editing strategy for improving performance of β-glucuronidase from Aspergillus terreus[J]. Chemical Engineering Science, 2017, 167:145-153. |
| 8 | WU S, CHAPPELL J. Metabolic engineering of natural products in plants; tools of the trade and challenges for the future[J]. Current Opinion in Biotechnology, 2008, 19(2): 145-152. |
| 9 | NICOLAOU K C, YANG Z, LIU J J, et al. Total synthesis of taxol[J]. Nature, 1994, 367(6464): 630-634. |
| 10 | ZHAO Y J, LV B, FENG X D, et al. Perspective on biotransformation and De Novo biosynthesis of licorice constituents[J]. Journal of Agricultural and Food Chemistry, 2017, 65(51): 11147-11156. |
| 11 | KIM H J, TURNER T L, JIN Y S. Combinatorial genetic perturbation to refine metabolic circuits for producing biofuels and biochemicals[J]. Biotechnology Advances, 2013, 31(6): 976-985. |
| 12 | PADDON C J, WESTFALL P J, PITERA D J, et al. High-level semi-synthetic production of the potent antimalarial artemisinin[J]. Nature, 2013, 496(7446): 528-532. |
| 13 | CAO H, CHEN X Q, JASSBI A R, et al. Microbial biotransformation of bioactive flavonoids[J]. Biotechnology Advances, 2015, 33(1): 214-223. |
| 14 | SCHLAGER S, DRAGER B. Exploiting plant alkaloids[J]. Current Opinion in Biotechnology, 2016, 37:155-164. |
| 15 | O’CONNOR S E, MARESH J J. Chemistry and biology of monoterpene indole alkaloid biosynthesis[J]. Natural Product Reports, 2006, 23(4): 532-547. |
| 16 | SATO F, INUI T, TAKEMURA T. Metabolic engineering in isoquinoline alkaloid biosynthesis[J]. Current Pharmaceutical Biotechnology, 2007, 8(4): 211-218. |
| 17 | ZHANG R H, LI C Y, WANG J, et al. Microbial production of small medicinal molecules and biologics: from nature to synthetic pathways[J]. Biotechnology Advances, 2018, 36(8): 2219-2231. |
| 18 | SEKI H, SAWAI S, OHYAMA K, et al. Triterpene functional genomics in licorice for identification of CYP72A154 involved in the biosynthesis of glycyrrhizin[J]. Plant Cell, 2011, 23(11): 4112-4123. |
| 19 | SEKI H, OHYAMA K, SAWAI S, et al. Licorice β-amyrin 11-oxidase, a cytochrome P450 with a key role in the biosynthesis of the triterpene sweetener glycyrrhizin[J]. Proceedings of the National Academy of Sciences, 2008, 105(37): 14204-14209. |
| 20 | DAI Z B, LIU Y, SUN Z T, et al. Identification of a novel cytochrome P450 enzyme that catalyzes the C-2α hydroxylation of pentacyclic triterpenoids and its application in yeast cell factories[J]. Metabolic Engineering, 2019, 51: 70-78. |
| 21 | HAGEL J M, FACCHINI P J. Dioxygenases catalyze the O-demethylation steps of morphine biosynthesis in opium poppy[J]. Nature Chemical Biology, 2010, 6(4): 273-275. |
| 22 | ALLEN R S, MILLGATE A G, CHITTY J A, et al. RNAi-mediated replacement of morphine with the nonnarcotic alkaloid reticuline in opium poppy[J]. Nature Biotechnology, 2004, 22(12): 1559-1566. |
| 23 | FARROW S C, HAGEL J M, BEAUDOIN G A, et al. Stereochemical inversion of (S)-reticuline by a cytochrome P450 fusion in opium poppy[J]. Nature Biotechnology, 2015, 11(9): 728-732. |
| 24 | MEDEMA M H, OSBOURN A. Computational genomic identification and functional reconstitution of plant natural product biosynthetic pathways[J]. Natural Product Reports, 2016, 33(8): 951-962. |
| 25 | FIELD B, OSBOURN A E. Metabolic diversification—independent assembly of operon-like gene clusters in different plants[J]. Science, 2008, 320(5875): 543-547. |
| 26 | FIELD B, A-S FISTON-LAVIER, KEMEN A, et al. Formation of plant metabolic gene clusters within dynamic chromosomal regions[J]. Proceedings of the National Academy of Sciences of the United States of America, 2011, 108(38): 16116-16121. |
| 27 | WINZER T, GAZDA V, HE Z, et al. A Papaver somniferum 10-gene cluster for synthesis of the anticancer alkaloid noscapine[J]. Science, 2012, 336(6089): 1704-1708. |
| 28 | BROWN S F, BRANFORD A J, MORAN W. On the use of artificial neural networks for the analysis of survival data[J]. IEEE Transactions On Neural Networks, 1997, 8(5): 1071-1077. |
| 29 | KAUTSAR S A, SUAREZ DURAN H G, BLIN K, et al. PlantiSMASH: automated identification, annotation and expression analysis of plant biosynthetic gene clusters[J]. Nucleic Acids Research, 2017, 45(W1): W55-W63. |
| 30 | WEBER T, BLIN K, DUDDELA S, et al. AntiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters[J]. Nucleic Acids Research, 2015, 43(1): W237-W243. |
| 31 | DENOEUD F, CARRETERO-PAULET L, DEREEPER A, et al. The coffee genome provides insight into the convergent evolution of caffeine biosynthesis[J]. Science, 2014, 345(6201): 1181-1184. |
| 32 | ZHU M, WANG C X, SUN W T, et al. Boosting 11-oxo-beta-amyrin and glycyrrhetinic acid synthesis in Saccharomyces cerevisiae via pairing novel oxidation and reduction system from legume plants[J]. Metabolic Engineering, 2018, 45: 43-50. |
| 33 | HWANG E I, KANEKO M, OHNISHI Y, et al. Production of plant-specific flavanones by Escherichia coli containing an artificial gene cluster[J]. Applied and Environmental Microbiology, 2003, 69(5): 2699-2706. |
| 34 | TRENCHARD I J, SIDDIQUI M S, THODEY K, et al. De novo production of the key branch point benzylisoquinoline alkaloid reticuline in yeast[J]. Metabolic Engineering, 2015, 31:74-83. |
| 35 | NAKAGAWA A, MINAMI H, KIM J S, et al. A bacterial platform for fermentative production of plant alkaloids[J]. Nature Communications, 2011, 2(1): 326. |
| 36 | SIDDIQUI M S, KATE T, ISIS T, et al. Advancing secondary metabolite biosynthesis in yeast with synthetic biology tools[J]. FEMS Yeast Research, 2012, 12(2): 144-170. |
| 37 | HWANG E I, KANEKO M, OHNISHI Y, et al. Production of plant-specific flavanones by Escherichia coli containing an artificial gene cluster[J]. Applied & Environmental Microbiology, 2003, 69(5): 2699-2706. |
| 38 | DELOACHE W C, RUSS Z N, NARCROSS L, et al. An enzyme-coupled biosensor enables (S)-reticuline production in yeast from glucose[J]. Nature Chemical Biology, 2015, 11(7): 465-471. |
| 39 | DIETRICH J A, YOSHIKUNI Y, FISHER K, et al. A novel semi-biosynthetic route for artemisinin production using engineered substrate-promiscuous P450BM3[J]. ACS Chemical Biology, 2009, 4(4): 261-267. |
| 40 | CHANG M C Y, EACHUS R A, TRIEU W, et al. Engineering Escherichia coli for production of functionalized terpenoids using plant P450s[J]. Nature Chemical Biology, 2007, 3(5): 274-277. |
| 41 | SUN W T, XUE H J, LIU H, et al. Controlling chemo- and regioselectivity of a plant P450 in yeast cell toward rare licorice triterpenoid biosynthesis[J]. ACS Catalysis, 2020,10: 4253-4260. |
| 42 | DELOACHE W C, RUSS Z N, NARCROSS L, et al. An enzyme-coupled biosensor enables (S)-reticuline production in yeast from glucose[J]. Nature Chemical Biology, 2015, 11(7): 465-471. |
| 43 | XIE W P, LV X M, YE L, et al. Construction of lycopene-overproducing Saccharomyces cerevisiae by combining directed evolution and metabolic engineering[J]. Metabolic Engineering, 2015, 30:69-78. |
| 44 | WANG P P, WEI W, YE W, et al. Synthesizing ginsenoside Rh2 in Saccharomyces cerevisiae cell factory at high-efficiency[J]. Cell Discovery, 2019, 5(1): 1-14. |
| 45 | EDGAR S, LI F S, QIAO K, et al. Engineering of taxadiene synthase for improved selectivity and yield of a key taxol biosynthetic intermediate[J]. ACS Synthetic Biology, 2016, 6(2): 201-205. |
| 46 | XIONG S T, WANG Y, YAO M D, et al. Cell foundry with high product specificity and catalytic activity for 21-deoxycortisol biotransformation[J]. Microbial Cell Factories, 2017, 16(1): 105. |
| 47 | GONZALEZ F J, KORZEKWA K R. Cytochromes P450 expression systems[J]. Annual Review of Pharmacology & Toxicology, 1995, 35(1): 369-390. |
| 48 | BIGGS B W, LIM C G, SAGLIANI K, et al. Overcoming heterologous protein interdependency to optimize P450-mediated Taxol precursor synthesis in Escherichia coli[J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(12): 3209-3214. |
| 49 | MUNRO A W, GIRVAN H M, MCLEAN K J. Variations on a (t)heme—novel mechanisms, redox partners and catalytic functions in the cytochrome P450 superfamily[J]. Natural Product Reports, 2007, 24(3): 585-609. |
| 50 | GALANIE S, THODEY K, TRENCHARD I J, et al. Complete biosynthesis of opioids in yeast[J]. Science, 2015, 349(6252): 1095-1100. |
| 51 | SOH K C, HATZIMANIKATIS V. Dreams of metabolism[J]. Trends in Biotechnology, 2010, 28(10): 501-508. |
| 52 | RENAULT H, BASSARD J E, HAMBERGER B, et al. Cytochrome P450-mediated metabolic engineering: current progress and future challenges[J]. Current Opinion in Plant Biology, 2014, 19: 27-34. |
| 53 | PADDON C J, KEASLING J D. Semi-synthetic artemisinin: a model for the use of synthetic biology in pharmaceutical development[J]. Nature Reviews Microbiology, 2014, 12(5): 355-367. |
| 54 | YANG D, PARK S Y, PARK Y S, et al. Metabolic engineering of Escherichia coli for natural product biosynthesis[J]. Trends in Biotechnology, 2020, 38(7):745-765. |
| 55 | ZHAO Y J, FAN J J, WANG C, et al. Enhancing oleanolic acid production in engineered Saccharomyces cerevisiae[J]. Bioresource Technology, 2018, 257:339-343. |
| 56 | LIAO P, HEMMERLIN A, BACH T J, et al. The potential of the mevalonate pathway for enhanced isoprenoid production[J]. Biotechnology Advances, 2016, 34(5): 697-713. |
| 57 | LIANG C N, ZHANG X X, WU J Y, et al. Dynamic control of toxic natural product biosynthesis by an artificial regulatory circuit[J]. Metabolic Engineering, 2020, 57: 239-246. |
| 58 | SCALCINATI G, KNUF C, PARTOW S, et al. Dynamic control of gene expression in Saccharomyces cerevisiae engineered for the production of plant sesquitepene alpha-santalene in a fed-batch mode[J]. Metabolic Engineering, 2012, 14(2): 91-103. |
| 59 | SKJOEDT M L, SNOEK T, KILDEGAARD K R, et al. Engineering prokaryotic transcriptional activators as metabolite biosensors in yeast[J]. Nature Chemical Biology, 2016, 12(11): 951-958. |
| 60 | SIEDLER S, STAHLHUT S G, MALLA S, et al. Novel biosensors based on flavonoid-responsive transcriptional regulators introduced into Escherichia coli[J]. Metabolic Engineering, 2014, 21: 2-8. |
| 61 | LIU D, XIAO Y, EVANS B S, et al. Negative feedback regulation of fatty acid production based on a malonyl-CoA sensor-actuator[J]. ACS Synthetic Biology, 2015, 4(2): 132-140. |
| 62 | XU P, WANG W Y, LI L Y, et al. Design and kinetic analysis of a hybrid promoter-regulator system for malonyl-CoA sensing in Escherichia coli[J]. ACS Chemical Biology, 2013, 9(2): 451-458. |
| 63 | LANGAN R A, BOYKEN S E, NG A H, et al. De novo design of bioactive protein switches[J]. Nature, 2019, 572:205-210. |
| 64 | ZHAO E M, ZHANG Y, MEHL J, et al. Optogenetic regulation of engineered cellular metabolism for microbial chemical production[J]. Nature, 2018, 555(7698): 683-687. |
| 65 | QIN J F, ZHOU Y J, KRIVORUCHKO A, et al. Modular pathway rewiring of Saccharomyces cerevisiae enables high-level production of L-ornithine[J]. Nature Communications, 2015, 6(1): 1-11. |
| 66 | ZHAO S, JONES J A, LACHANCE D M, et al. Improvement of catechin production in Escherichia coli through combinatorial metabolic engineering[J]. Metabolic Engineering, 2015, 28: 43-53. |
| 67 | GAIRIK S, ABHISHEK G, DAVID G, et al. In vivo co-localization of enzymes on RNA scaffolds increases metabolic production in a geometrically dependent manner[J]. Nucleic Acids Research, 2014, 42(14):9493-9503. |
| 68 | BROWN S, CLASTRE M, COURDAVAULT V, et al. De novo production of the plant-derived alkaloid strictosidine in yeast[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015, 112(11): 3205-3210. |
| 69 | DAI Z J, HUANG M T, CHEN Y, et al. Global rewiring of cellular metabolism renders Saccharomyces cerevisiae Crabtree negative[J]. Nature Communications, 2018, 9(1): 3059. |
| 70 | MITCHELL L A, WANG A, STRACQUADANIO G, et al. Synthesis, debugging, and effects of synthetic chromosome consolidation: synⅥ and beyond[J]. Science, 2017, 355(6329): eaaf4831. |
| 71 | SHEN Y, WANG Y, CHEN T, et al. Deep functional analysis of synⅡ, a 770-kilobase synthetic yeast chromosome[J]. Science, 2017, 355(6329):eaaf4791. |
| 72 | ZHOU K, QIAO K J, EDGAR S, et al. Distributing a metabolic pathway among a microbial consortium enhances production of natural products[J]. Nature Biotechnology, 2015, 33(4): 377-383. |
| 73 | JONES J A, VERNACCHIO V R, SINKOE A L, et al. Experimental and computational optimization of an Escherichia coli co-culture for the efficient production of flavonoids[J]. Metabolic Engineering, 2016, 35: 55-63. |
| 74 | JONES J A, VERNACCHIO V R, COLLINS S M, et al. Complete biosynthesis of anthocyanins using E. coli polycultures[J]. mBio, 2017, 8(3): e00621-e00617. |
| [1] | GAO Cong, CHEN Chenghu, CHEN Xiulai, LIU Liming. Progress and challenges of engineering microorganisms to produce biobased monomers [J]. Chemical Industry and Engineering Progress, 2023, 42(8): 4123-4135. |
| [2] | GUO Feng, ZHANG Shangjie, JIANG Yujia, JIANG Wankui, XIN Fengxue, ZHANG Wenming, JIANG Min. Biotransformation of one-carbon resources by yeast [J]. Chemical Industry and Engineering Progress, 2023, 42(1): 30-39. |
| [3] | TAO Yuxuan, ZHANG Shangjie, JING Yiwen, XIN Fengxue, DONG Weiliang, ZHOU Jie, JIANG Yujia, ZHANG Wenming, JIANG Min. Recent advances in the construction strategy of methylotrophic Escherichia coli [J]. Chemical Industry and Engineering Progress, 2021, 40(7): 3932-3941. |
| [4] | GUO Liang, GAO Cong, ZHANG Li, CHEN Xiulai, LIU Liming. Advances in the suitability of artificial metabolic pathways [J]. Chemical Industry and Engineering Progress, 2021, 40(3): 1252-1261. |
| [5] | WANG Ying, QU Junze, LIANG Nan, HAO He, YUAN Yingjin. Rapid construction and directed evolution of cell factories for carotenoid biosynthesis [J]. Chemical Industry and Engineering Progress, 2021, 40(3): 1187-1201. |
| [6] | LIU Weibing, YE Bangce. Progress of synthetic biology research and biological manufacturing of actinomycetes polyketides [J]. Chemical Industry and Engineering Progress, 2021, 40(3): 1226-1237. |
| [7] | MA Yueyuan, CHEN Jinchun, CHEN Guoqiang. Halophilic microorganisms as microbial chassis: applications and prospects [J]. Chemical Industry and Engineering Progress, 2021, 40(3): 1178-1186. |
| [8] | GAO Cong, GUO Liang, HU Guipeng, CHEN Xiulai, LIU Liming. Advances of metabolic flux regulation in microbial cell factories [J]. Chemical Industry and Engineering Progress, 2021, 40(12): 6807-6817. |
| [9] | Chen WANG, Meng ZHAO, Mingzhu DING, Ying WANG, Mingdong YAO, Wenhai XIAO. Application of biological scaffold system on synthetic biology [J]. Chemical Industry and Engineering Progress, 2020, 39(11): 4557-4567. |
| [10] | Pengcheng CHANG, Yang YU, Ying WANG, Chun LI. Combinatorial regulation strategies for efficient synthesis of terpenoids in Saccharomyces cerevisiae [J]. Chemical Industry and Engineering Progress, 2019, 38(01): 598-605. |
| [11] | YANG Kun, WANG Ying, LI Chun. Cell transporter protein and engineered applications [J]. Chemical Industry and Engineering Progress, 2017, 36(04): 1410-1417. |
| [12] | LIU Dingyu, MENG Jiao, WANG Zhiwen, CHEN Tao, ZHAO Xueming. Progress and application on multivariate modular metabolic engineering in metabolic engineering [J]. Chemical Industry and Engineering Progress, 2016, 35(11): 3619-3626. |
| [13] | XIAO Wenhai, ZHOU Sijie, WANG Ying, YUAN Yingjin. How to make biology more “engineering” [J]. Chemical Industry and Engineering Progree, 2016, 35(06): 1827-1836. |
| [14] | WAN Tao1,QI Haishan1,CHEN Yunlin2,WEN Jianping1. Research progress of biosynthesis of daptomycin and its derivative [J]. Chemical Industry and Engineering Progree, 2012, 31(07): 1581-1586. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||